Imagine that someone gives you a mystery novel with an entire page ripped out.
And let’s suppose someone else comes up with a computer program that reconstructs the missing page, by assembling sentences and paragraphs lifted from other places in the book.
Imagine that this computer program does such a beautiful job that most people can’t tell the page was ever missing.
DNA does that.
In the 1940’s, the eminent scientist Barbara McClintock damaged parts of the DNA in corn maize. To her amazement,
the plants could reconstruct the damaged section. They did so by copying other parts of the DNA strand, then pasting them into the damaged area.
This discovery was so radical at the time, hardly anyone believed her reports. (40 years later she won the Nobel Prize for this work.)
And we still wonder: How does a tiny cell possibly know how to do…. that???
A French HIV researcher and computer scientist has now found part of the answer. Hint: The instructions in DNA are not only linguistic, they’re beautifully mathematical. There is an Evolutionary Matrix that governs the structure of DNA.
Computers use something called a “checksum” to detect data errors. It turns out DNA uses checksums too. But DNA’s checksum is not only able to detect missing data; sometimes it can even calculate what’s missing. Here’s how it works.
In English, the letter E appears 12.7% of the time. The letter Z appears 0.7% of the time. The other letters fall somewhere in between. So it’s possible to detect data errors in English just by counting letters.
In DNA, some letters also appear a lot more often (like E in English) and some much less often. But… unlike English, how often each letters appears in DNA is controlled by an exact mathematical formula that is hidden within the genetic code table.
When cells replicate, they count the total number of letters in the DNA strand of the daughter cell. If the letter counts don’t match certain exact ratios, the cell knows that an error has been made. So it abandons the operation and kills the new cell.
Failure of this checksum mechanism causes birth defects and cancer.
Dr. Jean-Claude Perez started counting letters in DNA. He discovered that these ratios are highly mathematical and based on “Phi”, the Golden Ratio 1.618. This is a very special number, sort of like Pi. Perez’ discovery was published in the scientific journal Interdisciplinary Sciences / Computational Life Sciences in September 2010.

Jean-Claude Perez discovered an evolutionary mathematical matrix in DNA, based on the Golden Ratio 1.618
Before I tell you about it, allow me to explain just a little bit about the genetic code.
DNA has four symbols, T, C, A and G. These symbols are grouped into letters made from combinations of 3 symbols, called triplets. There are 4x4x4=64 possible combinations.
So the genetic alphabet has 64 letters. The 64 letters are used to write the instructions that make amino acids and proteins.
Perez somehow figured out that if he arranged the letters in DNA according to a T-C-A-G table, an interesting pattern appeared when he counted the letters.
He divided the table in half as you see below. He took single stranded DNA of the human genome, which has 1 billion triplets. He counted the population of each triplet in the DNA and put the total in each slot:
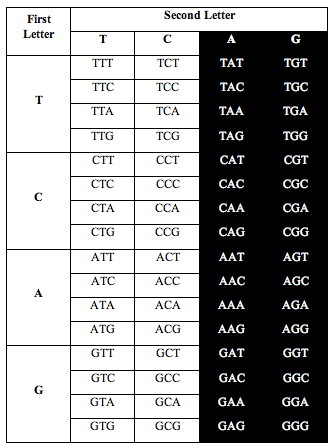
When he added up the letters, the ratio of total white letters to black letters was 1:1. And this turned out to not just be roughly true. It was exactly true, to better than one part in one thousand, i.e. 1.000:1.000.
Then Perez divided the table this way:

Perez discovered that the ratio of white letters to black letters is exactly 0.690983, which is (3-Phi)/2. Phi is the number 1.618, the “Golden Ratio.”
He also discovered the exact same ratio, 0.690983, when he divided the table the following two alternative ways:
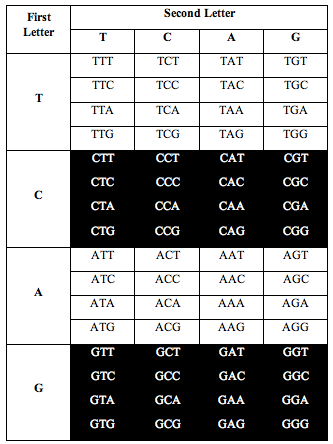
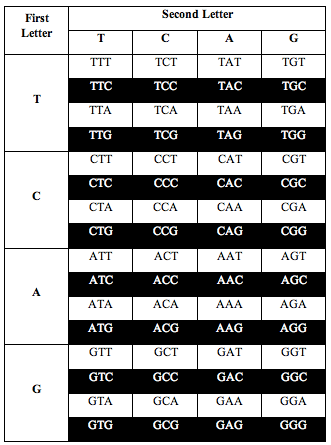
Again, the total number of white letters divided by the total number of black letters is 0.6909, to a precision of better than one part in 1,000.
Perez discovered two more symmetries:
 Above: Total ratio of white:black letters = 1:1
Above: Total ratio of white:black letters = 1:1
 Again, total ratio of white:black letters = 1:1
Again, total ratio of white:black letters = 1:1
So for three ways of dividing the table, the ratio of white to black is 1.000:1.000.
And for the other three ways of dividing it, the ratio is 0.690983 or (3-Phi)/2.
When you overlay these 6 symmetries on top of each other, you get a set of mathematical stairs with 32 golden steps. Then an absolutely fascinating geometrical pattern emerges: The “Dragon Curve” which is well known in fractal geometry. Here it is, labeled with DNA letters in descending frequency:
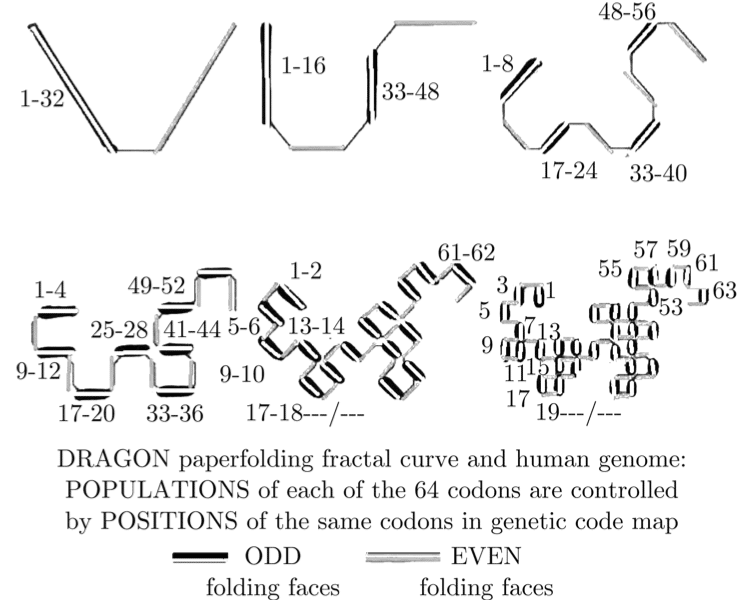

Animated Dragon Curve
You can see other non-DNA, computer generated versions of this same curve here.
Other interesting facts:
- Similar patterns with variations on these same rules are seen across a range of 20 different species. From the AIDS virus to bacteria, primates and humans
- Each character in DNA occurs a precise number of times, and each has a twin. TTT and AAA are twins and appear the most often; they’re the DNA equivalent of the letter E.
- This pattern creates a stair step of 32 frequencies, a specific frequency for each pair.
- The number of triplets that begin with a T is precisely the same as the number of triplets that begin with A (to within 0.1%).
- The number of triplets that begin with a C is precisely the same as the number of triplets that begin with G.
- The genetic code table is fractal – the same pattern repeats itself at every level. The micro scale controls conversion of triplets to amino acids, and it’s in every biology book. The macro scale, newly discovered by Dr. Perez, checks the integrity of the entire organism.
- Perez is also discovering additional patterns within the pattern.
I am only giving you the tip of the iceberg. There are other rules and layers of detail that I’m omitting for simplicity. Perez presses forward with his research; more papers are in the works, and if you’re able to read French, I recommend his book “Codex Biogenesis” and his French website. Here is an English translation.
(By the way, he found some of his most interesting data in what used to be called “Junk DNA.” It turns out to not be junk at all.)
OK, so what does all this mean?
- Copying errors cannot be the source of evolutionary progress, because if that were true, eventually all the letters would be equally probable.
- This proves that useful evolutionary mutations are not random. Instead, they are controlled by a precise Evolutionary Matrix to within 0.1%
- When organisms exchange DNA with each other through Horizontal Gene Transfer, the end result still obeys specific mathematical patterns
- DNA is able to re-create destroyed data by computing checksums in reverse – like calculating the missing contents of a page ripped out of a novel.
No man-made language has this kind of precise mathematical structure. DNA is a tightly woven, highly efficient language that follows extremely specific rules. Its alphabet, grammar and overall structure are ordered by a beautiful set of mathematical functions.
More interesting factoids:
The most common pair of letters (TTT and AAA) appears exactly 1/13X as often as all the letters combined – consistently, the genomes of humans and chimpanzees.
If you put the 32 most common triplets in Group 1 and the 32 least common triplets in Group 2, the ratio of letters in Group1:Group2 is exactly 2:1. And since triplet counts occur in symmetrical pairs (TTT-AAA, TAT-ATA, etc), you can group them into four groups of 16.
When you put those four triplet populations on a graph, you get the peace symbol:
Does this precise set of rules and symmetries appear random or accidental to you?
My friend, this is how it is possible for DNA to be a code that is self-repairing, self-correcting, self-re-writing and self-evolving. It reveals a level of engineering and sophistication that human engineers could only dream of. Most of all, it’s elegant.
Cancer has sometimes been described as “evolution run amok.” Dr. Perez has noted interesting distortions of this matrix in cancer cells. I strongly suspect that new breakthroughs in cancer research are hidden in this matrix.
I submit to you that the most productive research that can possibly be conducted in medicine and computer science is intensive study of the DNA Evolution Matrix. Like I said, this is just the tip of the iceberg.
There is so much more here to discover!
When we develop computer languages based on DNA language, they will be capable of extreme data compression, error correction, and yes, self-evolution. Imagine: Computer programs that add features and improve with time. All by themselves.
What would that be like?
Perry Marshall
P.S.: Dr. Perez and I are friends. Perez worked on HIV research with the man who originally discovered HIV, Luc Montagnier. Perez also worked in biomathematics and Artificial Intelligence at IBM. I’m familiar with this work because last spring I had the privilege of helping him translate his groundbreaking research paper about this into English.
You can read it here: “Codon Populations in Single-stranded Whole Human Genome DNA Are Fractal and Fine-tuned by the Golden Ratio 1.618”
Click here for a more in-depth PDF version of this report.
Download The First 3 Chapters of Evolution 2.0 For Free, Here – https://evo2.org/evolution/Where Did Life And The Genetic Code Come From? Can The Answer Build Superior AI? The #1 Mystery In Science Now Has A $10 Million Prize. Learn More About It, Here – https://www.herox.com/evolution2.0













From Risalei Nur collection by Said Nursi
Through the light of belief, man rises to the highest of the high and acquires a value worthy of Paradise. And through the darkness of unbelief, he descends to the lowest of the low and falls to a position fit for Hell. For belief connects man to the All-Glorious Maker; it is a relation. Thus, man acquires value by virtue of the Divine art and inscriptions of the dominical Names which become apparent in him through belief. Unbelief severs the relation, and due to that severance the dominical art is concealed. His value then is only in respect to the matter of his physical being. And since this matter has only a transitory, passing, temporary animal life, its value is virtually nothing.
Man is such an antique work of art of Almighty God. He is a most subtle and graceful miracle of His power whom He created to manifest all his Names and their inscriptions, in the form of a miniature specimen of the universe. If the light of belief enters his being, all the meaningful inscriptions on him may be read. As one who believes, he reads them consciously, and through that relation, causes others to read them. That is to say, the dominical art in man becomes apparent through meanings like, “I am the creature and artefact of the All-Glorious Maker. I manifest His mercy and munificence.” That is, belief, which consists of being connected to the Maker, makes apparent all the works of art in man. Man’s value is in accordance with that dominical art and by virtue of being a mirror to the Eternally Besought One. In this respect insignificant man becomes God’s addressee and a guest of the Sustainer worthy of Paradise superior to all other creatures.
However, should unbelief, which consists of the severance of the relation, enter man’s being, then all those meaningful inscriptions of the Divine Names are plunged into darkness and become illegible. For if the Maker is forgotten, the spiritual aspects which look to Him will not be comprehended, they will be as though reversed. The majority of those meaningful sublime arts and elevated inscriptions will be hidden. The remainder, those that may be seen with the eye, will be attributed to lowly causes, nature, and chance, and will become utterly devoid of value. While they are all brilliant diamonds, they become dull pieces of glass. His importance looks only to his animal, physical being. And as we said, the aim and fruit of his physical being is only to pass a brief and partial life as the most impotent, needy, and grieving of animals. Then it decays and departs. See how unbelief destroys human nature, and transforms it from diamonds into coal.
Dear Perry
I accept your algorithmic evolution theory but not the millions of years. Any reading of the Bible rules out millions and billions. You need to go back to Genesis. My investigations of the proofs for long ages lead me to believe that when all the evidence is considered including the fossil records that long ages are not at all certain. Why do you have so much invested in the long age ideas?
Perry, I did read your book. What is God’s role in the model of evolution that you propose? Is he just the designer of the laws of nature according to which life originates by chance and then evolves in an unguided manner? Do you believe that God interferes into his creation by e.g. originating the first life form?
Les,
I hesitate to say. On the face of it, based on what we know RIGHT NOW, the genetic code has every appearance of being designed. Especially to a communications engineer who used to work at a networking company, where people developed their own networking languages on an almost daily basis.
However I am wary of “God of Gaps” arguments. I talk about this in chapter 24. There is ALWAYS a gap, and if you understand Godel’s incompleteness theorem, there ALWAYS will be. It is fundamentally impossible for science to ever answer all the questions it raises. (Which is why scientism is a fool’s errand.) But increasingly I’m disinclined to point to any ONE single fact, or measurement, or phenomenon, and say “Here it is right here, THIS is the finger of God.” I’m more inclined to say it’s found in the totality and grandeur of nature as a whole.
A LOT of Christians are, whether they realize it or not, looking for that specific finger-of-God piece of data. It’s very comforting to find it… but then disquieting if someone then comes up with a natural explanation and pulls the rug out from under you. Newton thought that the proof of God was that the universe didn’t collapse from gravitational attraction. Well, now we know that laws account for that. I find that philosophically informed Christians are nervous about God of Gaps arguments.
What we don’t know is where the laws come from. That question always seems to point us back towards God.
What I’m comfortable with is that at the highest level, belief in God gives us grounds for believing that the universe is intelligible, discoverable, rational and mathematical. And that there’s always another layer of discovery. You also consistently find that, at the edges of science, atheists often revert to random chance, “the universe is senseless and incomprehensible,” “we may never solve this,” “there’s an infinite number of universes and this is just one of the lucky ones” and other anti-scientific, non-testable forms of abdication. Belief in God propels you through that muck. It worked 500 years ago and it still works today.
As for my own “Where is God in all this question” – I find the answer more readily in my personal experiences with God and in miracles. I’ve seen quite a few first hand, see http://www.coffeehousetheology.com/miracles.
I think it’s curious that many conservative evangelical Christians who embrace beliefs like Young Earth Creationism (a 6-day series of VERY LARGE miracles) tend to overwhelmingly as a group NOT believe in the gifts of the Holy Spirit.
It strikes me that their rejection of the latter (in my view their arguments for rejecting miracles are weak and unbiblical) is why they cling so tightly to YEC. Everyone, after all, gropes to find the presence of God in their life somewhere. I even find most atheists wanted that at one time – and then gave up.
ancient aliens
I’m afraid my conclusion after reading the article is that it strengthens the case for ‘old-school darwinism’. But I agree about the nonmathematical biologists’ lack of understanding of the subject. They are merely believers. But since evolution works they are not really to blame for adhering to an idea they dont fully grasp. I consider the problem of evolution by random mutation to be still unproven but since the disproof is absent time is on the side of the usual view. The problem with proving it is about outruling significant probability for divergent processes, which despite selection would lead to the end of life or to some other unfavorable overall condition.
The article simply shows that DNA regulates the extent of available mutation which eliminates the problem of strongly varying environmental conditions. DNA seems to be well-engineered whether or not it happened on purpose or by chance.
An entirely different aspect is that there is no profound theory of matter available. Quantum mechanics is an extremely economical recepy for extracting quantitative information about something, the nature of which isnt even believed to exist independently. So there is room for a great mystery beyond the chemical model of DNA, even though I dont think this article makes that apparent at all. Sorry 🙂
Old-school Neo-Darwinism is crumbling fast. See http://www.evo2.org/royal-society-evolution.
I read through your entire blog last night and this was my favorite. I am a civil/hydro engineer and spend most of my time creating mathematical models for various simulations. I appreciate your approach to applying information theory to DNA. I came across this article this morning and immediately thought back to reading this last night. The article title is “Physicists Confirm There’s a Second Layer of Information Hidden in Our DNA”
Theoretical physicists have confirmed that it’s not just the information coded in our DNA that shapes who we are—it’s also the way DNA folds itself that controls which genes are expressed inside our bodies.
here is the link, I hope it will be helpful in your work.
https://futurism.com/physicists-confirm-theres-a-second-layer-of-information-hidden-in-our-dna/
Hi Perry, I’ve just finished your fascinating book. Thank you so much for writing it!
I’m a Christian chemical and process engineer who has always wanted to be able to reconcile science with my faith, and I have found your book to be a very helpful stepping stone towards achieving that.
In the book you mention that DNA folds up in a fractal pattern to take up minimal space without tangling.
Then at the end of the book you mentioned very briefly about the very recent discovery of fractal patterns in the DNA sequence.
Your above post unpacks in much more detail what you meant by fractal patterns in the DNA, and it sounds like it is focused on overall patterns of frequency in occurrence, rather than patterns in the sequential ordering of the DNA triplets. But do you know whether there are also patterns in the ordering of DNA triplets along the DNA strand, and whether this might be what dictates the fractal folding pattern of the DNA?
Thanks for your work, it’s so enlightening!
Thanks for reading the book! I hope you’ll write us an Amazon review.
Your question brings up two issues which as far as I can tell are separate: The fractal pattern of codons, and the fractal globule for folding the physical helix (see https://hms.harvard.edu/news/dna-folding-neatniks-dream).
For the former, Jean-Claude Perez has written many papers about this and you can easily find them by searching.
So glad to be of help.
Prof Prem raj Pushpakaran writes — let us celebrate 23rd November as Fibonacci Day!!!
The conventional theory of evolution is alive and well, and more and more evidence will be added to perfect the picture. However just like physics, it is a theory which is based on a reference frame of a provisional character. The properties of the subatomic world are handled in a bookkeeping manner rather than to be deeply understood. There is no established theory of electric charge. It just is there.
Since electromagnetism is quantitatively covered by current theory the unused parts of its invariance group may hold the key to unlocking those only bookkept basic concepts. I have touched that in previous comments here. My theory is that the subatomic world as it appears to us is the result of a motional effect. Where approximately logarithmic space-time rules in what appears like a small scale to us.
In internal coordinates the force is falling off exponentially, like the strong interaction. If the falloff is transformed to labcoordinates the distance dependence is 1/r^2. Suggesting that the nuclear forces are motional transformations of 1/r^2 forces but if this is correct, subatomic physics would have to be completely rewritten to make sense of it. Further the 1/r2 forces too might as well be another aspect of the motional effects. The Coulomb force or maybe also gravity?More akin to Einsteins equivalence principle, where constant acceleration and constant gravity are seen as equivalent. Since the invariance group of electromagnetism in the case of spacetime inversions points to constant acceleration in contracting and expanding phases, there is a spatial turning point in between them. A minimum waist. If that turning point is synchronised with something which determines our reality, then maybe, I speculate, it is what we perceive as gravity. In any case it seems to be in accordance with the mentioned equivalence principle and ought to be worth considering.
One unfamiliar aspect of such a theory of gravity is that there is a length associated with it. For earths gravity the associated length is about a lightyear. We clearly dont see that associated with the earth. But if the distance is much bigger than it appears that might explain why gravity appears to be so much weaker
GRT doesnt cover this case even though it does cover the case of the full swing of that kind of motion. But then would appear to need negative mass to drive it as, maybe in the case of dark matter. But I am not considering the full swing, just a synchronised phase of it. The invariant type of motion is then seen as a part of a periodic phenomenon, only accessible in a specific phase. But this idea of bringing in gravity is an aside that I have been repelling mostly although it keeps coming back. For example the difference between the Coulomb force and gravity may be that the coulomb force, the charges represent more than a single lockpoint so we see two different turning points/lockpoints while with gravity we are locked to a single phase of the cycle.
And dark + ordinary matter might complete the relatedness with the Coulomb force.
Anyway.
The idea is that everything we perceive is phaselocked to an underlying pulsating phenomenon, which we cannot access other than through theory. This because we dont exist outside of the turning points or lockpoint(s). That pulsating phenomenon I intuitively connect both with consecutive contraction and expansion~ spacetime inversion as it appears in the invariance group of elelctromagnetism. And simultaneously I see it as maybe the way a number crunching cycle could appear at some level. In the extremes of that cycle there would be turning points, where everything came to a standstill. I am guessing that is our reality. A series of turning points at a higher level. Our continuous time then consists in discrete moments in the higher reality. But not necessarily possible to discern for us.
And the higher level time difference between turning points might be immense while taking up zero time for us.
Back to biology. The theory of evolution might be alive and well however we dont know why those molecules are so stable and persistent. I suggest that physicists dont even know whether physics can explain it. The calculations are simply too complicated.
If the physics theory I have been sketching is anyway near correct it means both that there may be immense timespans available at the higher level so the improbable could well be selected by suitable feedback mechanisms. Just like the ordinary theory of evolution employs multiple kinds of feedback.
The theory of intelligent design becomes more tractable but also fully compatible with conventional evolution.
Everything is about feedback. The logarithmic spacetime I mentioned combined with a periodic or quasiperiodic phenomenon(impossible to perceive by us) implies that there may be feedback. And probably nonlinearity of some kind, I like to think. Due to the potentially immense times available at the higher level there may be room for things beyond our imagination and yet fully compatible with a basic underlying randomness.
When logarithmic/linear/exponential time concepts are in operation the usual conclusions from thermodynamics and say, Poincare cycles, are altered. Poincare cycles may become something mundane.
Perry, another évidence of GENOME DNA CHECKSUM IS this HGO (Human Génome Optimum). Please read
https://www.google.com/url?sa=t&source=web&rct=j&url=https://crimsonpublishers.com/nacs/pdf/NACS.000512.pdf&ved=2ahUKEwigicSLoo7mAhVqBWMBHRBBAIkQFjABegQIAxAB&usg=AOvVaw1Of1k36onv_dGDtElSsEdp
I am very glad you are doing this work!
Perry’s suggested Human Génome CHECKSUM HGO Human Génome Optimum in other Cancers liké breast, prostate etc…
https://www.google.com/url?sa=t&source=web&rct=j&url=https://juniperpublishers.com/ctoij/pdf/CTOIJ.MS.ID.555756.pdf&ved=2ahUKEwih4P6eqo_mAhWC3eAKHZSPAl8QFjAAegQIBBAB&usg=AOvVaw0W66VesEZYHAvu4Jtxn-h2
Hi, I am aware that this particular blog page has been inactive for quite a while, but I hope that you still will be able to pick this up. I find the findings of Dr Perez very intriguing (but got across your webpage only today!). I do not read French and cannot read his book. Do you know if his table with frequences of different codon values is based on the whole genome, or only the protein-coding parts? Has he tried to replace the codon values with the amino acids they represent, to see if the corresponding counts display something of interest? If you can provide his contact details, I can also ask him myself 🙂
He has many papers published in English, you can find them all on Google Scholar.
It is of some interest that Leibniz, Cantor and Goedel all saw God behind science.
Goedel even ventured an onthological proof of Gods existence.
The reason they came to that conclusion is that they all encountered an intellectual challenge in moving beyond the reaches of an existing logical system, which necessitated a creative response.
Creativity implies our divine origin according to some thinkers.
Lyndon Larouche has often returned to that viewpoint and put it into contrast with Dawkins views.
Personally I see no harm in Dawkins arguments, since it offers a challenge for others to contribute. Dawkins offers a clear conceptual model.
I think its weak side is primarily that it is based on a very incomplete materialist basis.
The sciences are descriptive but dont explain how nature can be that way.
So no theory based on current science can be very deep.
The idea of connecting creativity to our divine origin is opposed to the thesis that we’re an aggregate of blind material processes.
Black-boxes, soul-less input-output systems.
Behaviourism is based on that latter idea.
While creativity unbinds our apparent frailty and instills hope for a greater future.
Historically german philosophers have beeen on both sides of the debate.
The meaning of the concept of God is something beyond our current understanding.
Something we might consider benevolent, in view of the many orderly and meaningful facets we encounter as we study the facts of the world.
But being beyond means we dont quite understand what it is.
Then what happens as science develops further and we begin to discern more and understand more about what we previously just summarised under the notion of the divine?
Will we then see a higher level separating us from the next version of the beyond? So God would now turn out to be higher up the ladder?
Or will there be an ending hierarchy?
Like the wizard of Oz turning out to be an old man behind a smokescreen?
The above philosophers, at least Cantor, probably didnt think there would be a finite ladder. After all he saw an infinite succession of ordinals, the alefs whereas God was seen as an absolute.
In my view Darwins idea is useful as a filter for sorting out the best solutions, given some creative input. So it doesnt really conflict with the idea of God.
God would likely apply it to perfect creation.
But to take a different perspective:
Seeing math in DNA, isnt incompatible with the idea that the world has emerged from material causes. Some kind of nonlinear feedback arose on the way and the universe became capable of creating complex codes representing everything, living and dead matter.
And that at some point consciousness resulted: a feedback phenomenon. Selfreference.
The universe became conscious.
God emerged out of the chaos.
Biological systems depend on highly advanced machinery. All of which is fabricated by perfect components. What industry is capable of massproducing with that precision?
Answer: A computer code has that degree of precision.
So the computer paradigm is very tempting to apply in a theory of everything.
Some scientists suspect the Universe really is a computer simulation.
Call it a simulation or call it a conscious universe based on an undertlying mathematical framework.
Where’s the difference between the idea that God always was and the idea that God emerged through various nonlinear processes?
(My pet idea is that there are different time concepts through which probability theory opens the opportunity to render the likelihood of emerging intelligence approach 100% in some of the related reference systems)
This is a long thread and I have posted some comments before and along the way I have returned to the following conclusion:
Seems to me it is more likely that both the materialist and the a priori religious worldview are fully compatible when the big picture is rendered more discernable and that the two opposing camps may arrive at a consensus.
Thank you for your thoughtful comments.